Search

体外转录概述 |
在实验室中合成 RNA 对于许多技术至关重要,包括分析、生化和分子生物学研究。体外转录(IVT)是一种简单的方法,可以实现任何序列的 RNA 分子的模板指导合成——从短链寡核苷酸到数千碱基对,产量从微克到毫克不等。IVT mRNA 的生成涉及在目标序列上游设计带有噬菌体启动子的模板,然后使用相应的 RNA 聚合酶进行转录。本综合概述将介绍:
体外转录简介
通过使用反应组分,如缓冲系统、核糖核苷酸三磷酸(NTPs)、DNA模板和噬菌体RNA聚合酶,研究人员可以高效地制备高质量的RNA。本概述将带您了解IVT的关键方面,包括所需组分、合成流程、递送方法、规模化技术及多种应用,为您全面理解这一重要的科研工具。

体外转录反应所需组分
体外转录需要一个纯化的线性DNA模板(包含启动子)、核糖核苷三磷酸(NTPs)、包含DTT和镁离子的缓冲体系,以及合适的噬菌体RNA聚合酶来合成mRNA。转录反应所用的具体条件取决于特定应用所需的RNA量。
缓冲体系
体外转录(IVT)中的缓冲体系旨在为转录反应创造最佳环境。将缓冲体系加入IVT反应的作用包括:
- 维持最佳pH值:缓冲液有助于确保pH值保持在有利于RNA聚合酶活性的范围内。
- 提供必需的离子:如镁离子(Mg²⁺)和钾离子(K⁺)等离子对于转录过程中酶的活性和稳定性至关重要。
- 支持离子强度:维持反应混合物的整体离子强度对于关键酶的正确折叠和功能非常重要。
核糖核苷三磷酸(NTPs)
核糖核苷三磷酸(NTPs)对于转录过程至关重要,影响mRNA产量的效率和质量,因为它们是:
- RNA合成的构建模块:NTPs(ATP、CTP、GTP 和 UTP)是RNA聚合酶用于合成RNA链的必需底物。它们被掺入到不断延长的RNA链中,帮助确保根据DNA模板形成正确的序列。
- 能量来源:NTP中的高能磷酸键为聚合反应提供所需的能量。最佳的NTP浓度可确保高效的转录过程。
- 产量和质量的决定因素:NTP的浓度直接影响所合成RNA的产量和质量。
DNA模板
在体外转录(IVT)反应中,DNA模板通过以下关键功能为RNA分子的合成提供序列信息指导:
- 序列蓝图:DNA模板包含将被转录为RNA的核苷酸序列。RNA聚合酶读取DNA模板,并合成与DNA序列互补的RNA链。
- 启动子识别:DNA模板通常包含一个启动子区域,这是RNA聚合酶能够识别的特定序列。该启动子对于在DNA模板的正确位置启动转录至关重要。
- 方向性:DNA模板决定了转录的方向。RNA聚合酶沿着DNA模板链以3'到5'的方向移动,同时以5'到3'的方向合成RNA链。
Jump to IVT Templates to learn more about plasmid, PCR, oligonucleotide, and cDNA templates.
噬菌体RNA聚合酶
RNA聚合酶是一类将DNA序列复制为RNA序列的酶,在IVT反应的起始和延伸过程中发挥作用。所有真核生物都拥有不同的聚合酶,用于转录不同类型的基因。一些类型包括T7、T3和SP6:
- T7 RNA聚合酶能够快速且选择性地从其自身启动子进行转录,合成速度更快,终止较少,优于*大肠杆菌*RNA聚合酶。
- T3 RNA聚合酶在结构上类似于T7,但由于特定氨基酸的替换导致与启动子的相互作用不同,因此在功能上有所区别。
- SP6 RNA聚合酶是一种稳定且高度特异性的酶,只能从其自身启动子进行转录,无法识别T3或T7 DNA。
IVT模板
为了促进启动子的识别,*体外*转录的DNA模板必须包含正确方向的双链RNA聚合酶启动子区域。这些模板包括质粒、PCR产物、寡核苷酸和cDNA:
许多常用的质粒克隆载体都包含噬菌体聚合酶启动子。它们通常在多克隆位点的两侧各有一个不同的启动子。质粒载体应通过限制性内切酶消化进行线性化。
可以通过在正向或反向PCR引物的5'端加入启动子序列,将启动子添加到PCR产物中。
将含有噬菌体启动子序列的两条互补寡核苷酸简单退火,即可制备双链DNA模板。
通过在逆转录过程中使用Invitrogen的oligo(dT)-T7启动子引物,可以从RNA生成转录模板。
模板的选择取决于具体应用、所需RNA的量,以及是否需要正义或反义转录本。
- 反义转录本:用于与mRNA杂交的探针(例如,Northern杂交、原位杂交)。
- 正义转录本:用于表达、结构或功能研究。
The +1 G of the RNA polymerase promoter sequence is the first base incorporated into the transcription product. For sense RNA, the 5' end of the coding strand must be adjacent to or just downstream of the +1 G. For antisense RNA, the 5' end of the noncoding strand must be adjacent to the +1 G.
合成IVT mRNA的关键步骤
在体外转录过程中发生的反应步骤,包括纯化、加帽、转录标记和加聚腺苷酸尾,对提高mRNA的稳定性、翻译效率和整体质量起着至关重要的作用。
纯化
用于分离聚腺苷酸(poly-A)RNA。POROS™ Oligo (dT)25亲和树脂是其中的优质产品。
能高效沉淀长度至少为100个核苷酸的RNA分子。
对乙醇沉淀小分子和大分子RNA都很有效,但不会沉淀核苷酸。
首选方法,利用硅胶树脂便于结合、洗涤和洗脱。
加帽
加帽可以保护mRNA免受核酸酶攻击,并促进其功能。
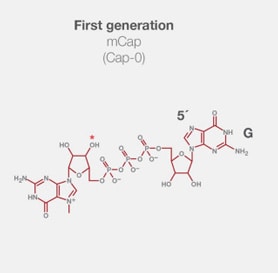
1. mCap类似物:由于可能出现反向取向,只有50%的加帽转录本可被翻译。

2. 反向帽类似物(ARCA):可防止反向取向,所有加帽转录本均可被翻译。

3. 三核苷酸加帽:产量和加帽效率更高(如CleanCap™)。可生成cap 1结构。
共转录加帽与转录后加帽
| 共转录加帽 | 转录后加帽 | |
|---|---|---|
| 过程
| 在转录过程中掺入帽模拟物 | 在转录反应后通过酶进行加帽 |
| 机制 | RNA聚合酶在5′端掺入帽结构 | cap0的酶促形成包括三个连续反应 |
| 加帽酶 | 帽结构在RNA出口通道周围结合 | 来自真核生物或病毒的酶 |
| 步骤 | 转录过程中一步完成 | 多步骤(水解、偶联、甲基化) |
| 效率 | 理论效率小于100% | 高效率 |
| 纯化 | 所需步骤更少 | 需要去除未使用的NTP |
| 应用 | 无缝RNA保护 | 可在体外产生加帽RNA |
转录标记
在转录过程中将标记核苷酸(荧光或放射性)掺入RNA,用于追踪定位或研究相互作用。
Poly-A尾的添加与翻译
Poly-A尾是在3'端非模板性地添加腺苷,在翻译和稳定性中起关键作用。
递送方法
转染是将核酸引入真核细胞的过程。mRNA被直接递送并在细胞质中表达。
脂质纳米颗粒(LNPs)
LNPs是适用于体内mRNA递送的载体。它们由四种成分组成:
- 可离子化脂质:与带负电荷的mRNA结合。
- 聚乙二醇化脂质:稳定颗粒。
- 磷脂:有助于结构形成。
- 胆固醇:有助于结构形成。
Researchers have found that lipid nanoparticle-mRNA formulations based on zwitterionic ionizable lipids can escape the endosome, leading to efficient protein expression and genome editing in vivo. Also, in addition to functioning as a delivery component, lipids can have therapeutic effects synergistic with mRNA-encoded proteins.[27] LNPs are composed of a few helper lipids which come in many shapes—for example, cylindrical-shaped lipid phosphatidylcholine can provide greater bilayer stability, which is important for in vivo application of LNPs.[28]
MESSENGER Max reagent, a Thermo Fisher Scientific product, significantly enhances the transcription process. It boasts up to five times the efficiency of DNA reagents in neurons and primary cells, transfecting more mRNA into these challenging cell types without the need for electroporation or viruses. This reagent offers over a two-fold improvement in transfection efficiency compared to other lipid-based options, faster protein expression without genomic integration risks, and up to ten times higher cleavage efficiency using mRNA CRISPRs.
Also see: mRNA Delivery Technology |Selecting a RNAi Strategy |Guidelines for RNA Transfection
扩大量体外转录(IVT)反应
- 常规:通常用于放射性标记的 RNA 探针(<100 微克)。
- 大规模:可生成 >100 微克 RNA(每微克模板可达 120–180 微克)。MEGAscript™ 技术允许在高核苷酸浓度下保持活性。
Conventional reactions
Conventional reaction conditions, such as those used in the Invitrogen MAXIscript Kit, use relatively low nucleotide concentrations (0.5 mM each). Higher nucleotide concentrations are not necessary since, in these reactions, the low concentration of radiolabeled or modified nucleotide present effectively limits the total yield of the reaction.
The total concentration of the limiting nucleotide (labeled/modified and unlabeled) should be at least 3 µM for efficient synthesis of full-length RNA transcripts of <400 nt (more will be needed to synthesize longer transcripts).
A 3 µM concentration of radiolabeled rNTP can be obtained by adding 5 µL of a 800 Ci/mmol, 10 mCi/mL (or 12.5 µM) solution of [α-32P] NTP. Higher specific activity labeled rNTPs are available but are provided at a much lower stock molar concentration (e.g. the 3000 Ci/mmol, 10 mCi/mL has a stock concentration of only 3.3 µM). Without the addition of unlabeled NTP, it is impossible to achieve the final minimum 3 µM reaction concentration.
Because limiting nucleotide concentration can result in premature termination of transcription, there is a trade-off between synthesis of high specific activity (or extensively modified) transcripts and full-length transcripts. Diluting the limiting radiolabeled or modified nucleotide with unlabeled nucleotide proportionally lowers the specific activity (or extent of modification) of the transcript but yields more full-length transcript. To make very high specific activity or extensively modified transcripts one should limit or omit any unlabeled limiting nucleotide present.
Large-scale synthesis
Large-scale in vitro transcription reactions can produce up to 120–180 µg RNA per microgram template in a 20 µL reaction. Invitrogen MEGAscript technology allows the phage RNA polymerases to remain active at high nucleotide concentrations that would ordinarily inhibit the enzyme. Yields from these large-scale reactions are typically 10 to 50 times higher than those possible with conventional transcription reactions (without any limiting nucleotide). Reaction conditions (e.g., the type of nucleotide salt, type and concentration of salt in the transcription buffer, enzyme concentration and pH) are all optimized not only for each polymerase but for the entire set of components. Only under these conditions can you achieve optimal yields.
IVT mRNA 试剂盒
Order IVT mRNA kits through Thermo Fisher Scientific. Most IVT mRNA kits and products above are available in multiple options based on your anticipated number of reactions.
按应用进行体外转录实验
| 反应规模 | 应用 | 技术/方法 |
|---|---|---|
| 常规/小规模 | 探针生成与RNA修饰 |
|
| 大规模 | aRNA扩增 |
|
| 表达研究 |
| |
| 结构分析 |
| |
| 机制研究 |
|
参考文献
- Beckert B, Masquida B. 体外转录合成RNA。2011年。《分子生物学方法》,703卷,第29–41页。
- Avci-Adali M, Behring A, Steinle H 等。体外合成修饰mRNA以诱导人细胞蛋白表达。2014年。《JoVE》。
- Pellestor F, Paulasova P. 肽核酸(PNA)。2004年。《欧洲人类遗传学杂志》。
- Muttach F, Muthmann N, Rentmeister A. 合成mRNA加帽。2017年。《贝尔斯坦有机化学杂志》。
- Muttach F, Muthmann N, Rentmeister A. Synthetic mRNA capping. 2017. Beilstein Journal of Organic Chemistry, 13, p. 2819–2832. doi: 10.3762/bjoc.13.274.
- Ramanathan A, Robb GB, Chan SH. mRNA加帽:生物学功能及应用。2016年。《核酸研究》。
- Martin SA, Moss B. Modification of RNA by mRNA guanylyl transferase and mRNA (guanine-7-) methyltransferase from vaccinia virions. 1975. Journal of Biological Chemistry, 250(24), p. 9330–9335.
- Shuman S. Structure, mechanism, and evolution of the mRNA capping apparatus. 2001. Process in Nucleic Acid Research and Molecular Biology, 66, p. 1–40. doi: 10.1016/s0079-6603(00)66025-7.
- Kore AR, Shanmugasundaram M, Charles I, Cheng AM, Barta TJ. Synthesis and application of 2’-fluoro substituted cap analogs. 2007. Bioorganic & Medicinal Chemistry Letters, 17(19), p. 5295–5299. doi: 10.1016/j.bmcl.2007.08.029.
- Jimielity J, Fowler T, Zuberek J, Stepinski J, Lewdorowicz M, Niedzwiecka A, Stolarski R, Darzynkiewics E, Rhoads RE. Novel “anti-reverse” cap analogs with superior translational properties. 2003. RNA, 9(9), p. 1108–1122. doi: 10.1261/rna.5430403.
- Shanmugasundaram M, Senthilvelan A, Kore AR. Recent Advances in Modified Cap Analogs: Synthesis, Biochemical Properties, and mRNA Based Vaccines. 2022. The Chemical Record, 22(8). doi: 10.1002/tcr.202200005.
- Nicholson AL, Pasquinelli AE. Tales of Detailed Poly(A) Tails. 2018. Trends in Cell Biology, 29(3), p. 191–200. doi: 10.1016/j.tcb.2018.11.002.
- Werner M, Purta E, Kaminska KH, Cymerman IA, Campbell DA, Mittra B, Zamudio JR, Sturm NR, Jaworski J, Bujnicki JM. 2'-O-ribose methylation of cap2 in human: function and evolution in a horizontally mobile family. 2011. Nucleic Acids Research, 39(11), p. 4756–4768. doi: 10.1093/nar/gkr038.
- Chong ZX, Yeap SW, Ho WY. Transfection types, methods, and strategies: a technical review. 2021. PeerJ, 9. doi: 10.7717/peerj.11165.
- Tabor S. Expression using the T7 RNA polymerase/promoter system. 2001. Current Protocols in Molecular Biology, 16(16.2). doi: 10.1002/0471142727.mb1602s11.
- Johannsen S, Paulus S, Dupre N, Muller J, Sigel RKO. Using in vitro transcription to construct scaffolds for one-dimensional arrays of mercuric ions. 2008. Journal of Inorganic Biochemistry, 102(5–6), p. 1141–1151. doi: 10.1016/j.jinorgbio.2007.12.023.
- Franzolin E, Pontarin G, Rampazzo C, Miazzi C, Ferraro P, Palumbo E, Reichard P, Bianchi V. The deoxynucleotide triphosphohydrolase SAMHD1 is a major regulator of SNA precursor pools in mammalian cells. 2013. PNAS, 110(35), p. 14272–14277. doi: 10.1073/pnas.1312033110.
- Miki K, Endo K, Takasashi S, Funakoshi S, Takei I, Katayama S, Toyoda T, Kotaka M, Takaki T, Umeda M, Okubo C, Nishikawa M, Oishi A, Narita M, Miyashiti I, Asano K, Hayashi K, Osafune K, Yamanaka S, Saito H, Yoshida Y. Efficient Detection and Purification of Cell Populations Using Synthetic MicroRNA Switches. 2015. Cell Stem Cell, 16(6), p. 699–711. doi: 10.1016/j.stem.2015.04.005.
- Bantle JA, Maxwell IH, Hahn WE. Specificity of oligo(dT)-cellulose chromatography in the isolation of polyadenylated RNA. 1976. Analytical Biochemistry, 72(1–2), p. 413–427. doi: 10.1016/0003-2697(76)90549-2.
- Cathala G, Savouret JF, Mendez B. A method for isolation of intact, translationally active ribonucleic acid. 1983. DNA, 2, p. 329–335.
- Walker SE, Lorsch J. Chapter Nineteen – RNA Purification – Precipitation Methods. 2013. Methods in Enzymology, 530, p. 337-343. doi: 10.1016/B978-0-12-420037-1.00019-1.
- Shi R, Lewis RS, Panthee DR. Filter paper-based spin column method for cost-efficient DNA or RNA purification. 2018. PloS One, 13(12). doi: 10.1371/journal.pone.0203011.
- Carter S. Introduction to Transcription. Chapter 9.2. Lumen Learning.
- Butler ET, Chamberlin MJ. Bacteriophage SP6-specific RNA polymerase. I. Isolation and characterization of the enzyme. 1982. Journal of Biological Chemistry, 257(10), p. 5772–5778.
- Pardi N, Tuyishime S, Muramatsu H, Kariko K, Mui BL, Tam YK, Madden TD, Hope MJ, Weissman D. Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. J Control Release 217, 345–351 (2015). doi: 10.1016/j.jconrel.2015.08.007.
- Cross R. Without these lipid shells, there would be no mRNA vaccines for COVID-19. 2021. Chemical and Engineering News, 8(99).
- Hou X, Zaks T, Langer R, Dong Y. Lipid nanoparticles for mRNA delivery. 2021. Nature Reviews Materials, 6, p. 1078–1094. doi: 10.1038/s41578-021–00358-0.
- Cheng X, Lee RJ. The role of helper lipids in lipid nanoparticles (LNPs) designed for oligonucleotide delivery. 2016. Advanced Drug Delivery Reviews, 99 Part A, p. 129–137. doi: 10.1016/j.addr.2016.01.022.
- Ling M-L, Risman SS, Klement JF, McGraw N, McAllister WT. Abortive initiation by bacteriophage T3 and T7 polymerases under conditions of limiting substrate. Nucl. Acids Res. (1989) 17: 1605–1618.
仅供科研使用,不可用于诊断目的。
