Search

Silencer Select siRNAs |
Achieve excellent gene knockdown with Silencer Select siRNAs
By using our Silencer Select siRNA, you can achieve clean, consistent phenotypic data, helping lead to more accurate and reproducible experimental outcomes. Our Silencer Select siRNAs are:
- Reliable—Benefit from our algorithm designs in the consistency and reliability of your phenotypic results.
- Precise—Our Locked Nucleic Acid (LNA) chemical modifications reduce off-target effects by up to 90% for greater specificity.
- Potent—Experience up to 100-fold greater potency compared to currently available siRNAs.
- Guaranteed—Silencer Select siRNA comes with a 100% guarantee to silence the target gene, offering one of the best guarantees in the industry.
Interested in targeting long non-coding RNAs?
Explore Silencer Select siRNAs for lncRNA or search below for your lncRNA gene of interest.
Targeted gene silencing with Silencer Select siRNAs
Click on the tabs below to view data on targeted gene silencing with Silencer Select siRNAs.
The Silencer Select design algorithm
At Thermo Fisher Scientific, we combined a powerful machine learning method with performance data from thousands of siRNAs to better understand the link between the sequence of an siRNA and its thermodynamic properties, target location, and silencing efficiency [1]. This is unlike most siRNA design algorithms which predict effective siRNAs that induce 70% target mRNA knockdown with only ~80% confidence and are inadequate for predicting more efficient siRNAs. RNAi applications demand better efficiency, leading us to build the Silencer Select algorithm and validate its functionality.
Silencer Select algorithm verification demonstrated our algorithm has:
- Increased predictive accuracy by 28% over the previous generation siRNA design algorithm (Figure 1)
- siRNAs designs that are up to 100-fold more potent than both modified and unmodified siRNAs from other suppliers
- siRNA designs with significantly higher percentage of “on-target” phenotypes compared to other siRNAs (Figure 2)
Figure 1. Silencer Select siRNA design algorithm significantly improves effective siRNA prediction accuracy. Silencer Select siRNAs consistently outperformed those designed with a previous algorithm, delivering stronger mRNA knockdown across 40 gene targets in HeLa cells. The inset shows percentage of siRNAs that achieved ≥70% and ≥80% knockdown, highlighting improved potency and design accuracy.
Silencer Select siRNAs enable our highest percentage of silenced phenotypes and benefit from being:
- More reliable in eliciting maximum knockdown levels
- More consistent in achieving the threshold level of knockdown required to see a loss-of-function phenotype
- More effective in producing the expected, silenced phenotypes compared to siRNAs from other vendors (Figure 2)

Enhanced specificity to reduce off-target effects
Sequence-specific off-target effects are one of the primary reasons for false positive results in RNAi experiments. In addition to the potency improvements afforded by the Silencer Select algorithm, advanced bioinformatic filtering criteria removes off-targeting due to sequence. Combined with incorporation of novel chemical modifications demonstrated to improve siRNA specificity, Silencer Select siRNAs are highly potent with low off-target effects.
Bioinformatic filters applied in the Silencer Select algorithm
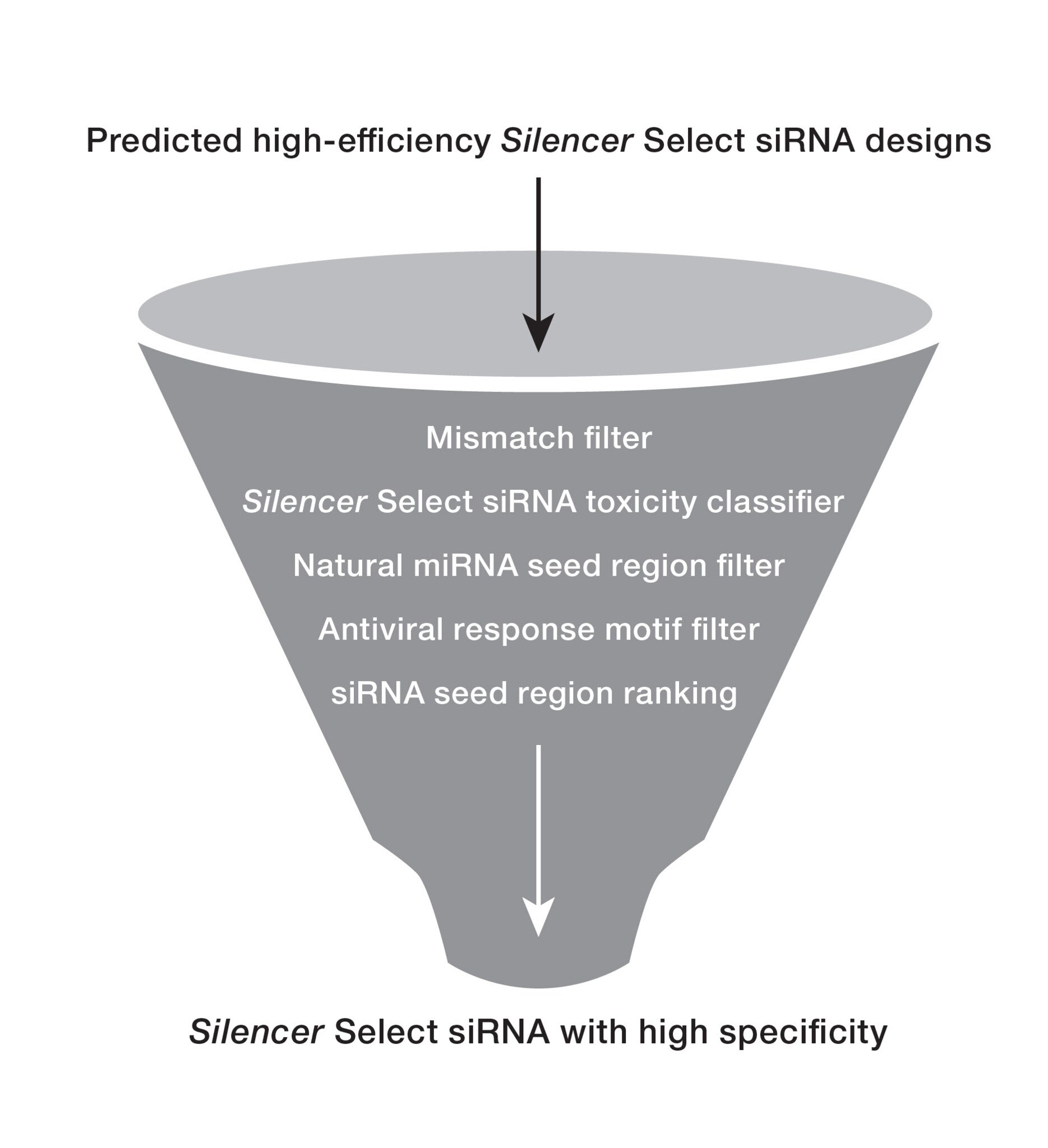
Although using siRNAs at low concentrations decreases off-target effects, further specificity gains can be made using bioinformatic filtering to predict and eliminate potentially inefficient siRNAs.
Our Silencer Select siRNA design process helps minimize off-target effects via:
- Stringent screening: identify and remove siRNAs with a high propensity for off-target effects by using a rigorous five-step process.
- Silencer Select toxicity classifier: helps eliminate sequences predicted to cause off-target apoptotic phenotypes to help ensure safer and more reliable siRNA performance.
- Seed region filtering: remove siRNAs with seed regions that resemble naturally occurring miRNAs and select those with the fewest seed region matches in the 3'UTRs of off-target transcripts to reduce miRNA pathway-related off-target effects.
The chemical modifications in Silencer Select siRNAs
Silencer Select siRNAs incorporate Locked Nucleic Acid (LNA) modifications to enhance their stability, binding affinity, and specificity. LNA-modified bases have the ribose ring chemically "locked" by a methylene bridge connecting the 2'-oxygen and the 4'-carbon. Adding LNA modifications to Silencer Select siRNA leads to:
- Consistent guide strand bias, which has been shown to correlate strongly with knockdown efficiency
- Prevention of the passenger strand from inducing silencing, which serves to reduce off-target effects (Figure 3)
- A reduced number of non-targeted, differentially expressed genes detected by gene expression array by up to 90% as compared to unmodified siRNAs (Figure 4)
- No negative impact on silencing efficiency and therefore do not compromise the expected on-target phenotypes

Figure 3. Silencer Select siRNA modifications reduce off-target effects and yield more reliable phenotypic data. Modified siRNAs preserved expected mitosis and apoptosis phenotypes for PLK and WEE1. In contrast the off-target apoptotic phenotypes elicited by 10 unmodified siRNAs were eliminated with addition of the Silencer Select modifications-demonstrating the modifications support generation of more reliable data.

Cleaner, more consistent data
siRNA mediated gene silencing enables the study of gene function with exceptional power and ease. However, currently available siRNAs for the same target do not always produce the same phenotype due to a combination of inconsistent silencing and sequence-specific off-target effects. Silencer Select siRNAs are designed with modifications that help overcome these issues, leading to more reliable data

Multiple publications confirm that higher siRNA concentrations lead to increased off-target effects [2,3]. This insight drove our efforts to design more potent siRNAs that can be used at lower concentrations, by significantly enhancing our siRNA design algorithms.
Silencer Select siRNAs
- Are up to 100X more potent than competitor siRNAs.
- Can be routinely transfected at concentrations of ≤5 nM and still retain their silencing power.
- Result in fewer off-target effects when used at these lower concentrations.
- Cost less per experiment than siRNAs used at higher concentrations.

Figure 5.Silencer Select siRNAs offer up to 100X higher potency compared to other siRNAs. Comparison of target knockdown in HeLa cells using Silencer Select siRNAs and siRNAs from two other suppliers to the same 10 different targets demonstrates the superior potency of Silencer Select siRNAs.
100% guarantee: One of the best guarantees in the industry
At Thermo Fisher Scientific, we guarantee that when you purchase two Invitrogen Silencer Select Pre-designed siRNA to the same target, those two siRNAs will silence the target mRNA by 70% or more. To qualify for the guarantee, siRNAs must have been transfected at ≥5 nM with proper controls and mRNA levels detected 48 hours post-transfection. Follow the link to learn more about our industry-leading guarantee.
Silencer Select validated siRNAs
A number of Invitrogen siRNAs targeting common human targets have been functionally tested by Thermo Fisher Scientific scientists and verified to reduce target mRNA levels by 70% or greater. The data sheet included with each of these siRNAs shows the extent of mRNA knockdown observed during testing and the exon targeted by the siRNA. We offer full sequences for predesigned and validated siRNAs with purchase.
The Silencer Select design algorithm
At Thermo Fisher Scientific, we combined a powerful machine learning method with performance data from thousands of siRNAs to better understand the link between the sequence of an siRNA and its thermodynamic properties, target location, and silencing efficiency [1]. This is unlike most siRNA design algorithms which predict effective siRNAs that induce 70% target mRNA knockdown with only ~80% confidence and are inadequate for predicting more efficient siRNAs. RNAi applications demand better efficiency, leading us to build the Silencer Select algorithm and validate its functionality.
Silencer Select algorithm verification demonstrated our algorithm has:
- Increased predictive accuracy by 28% over the previous generation siRNA design algorithm (Figure 1)
- siRNAs designs that are up to 100-fold more potent than both modified and unmodified siRNAs from other suppliers
- siRNA designs with significantly higher percentage of “on-target” phenotypes compared to other siRNAs (Figure 2)
Figure 1. Silencer Select siRNA design algorithm significantly improves effective siRNA prediction accuracy. Silencer Select siRNAs consistently outperformed those designed with a previous algorithm, delivering stronger mRNA knockdown across 40 gene targets in HeLa cells. The inset shows percentage of siRNAs that achieved ≥70% and ≥80% knockdown, highlighting improved potency and design accuracy.
Silencer Select siRNAs enable our highest percentage of silenced phenotypes and benefit from being:
- More reliable in eliciting maximum knockdown levels
- More consistent in achieving the threshold level of knockdown required to see a loss-of-function phenotype
- More effective in producing the expected, silenced phenotypes compared to siRNAs from other vendors (Figure 2)

Enhanced specificity to reduce off-target effects
Sequence-specific off-target effects are one of the primary reasons for false positive results in RNAi experiments. In addition to the potency improvements afforded by the Silencer Select algorithm, advanced bioinformatic filtering criteria removes off-targeting due to sequence. Combined with incorporation of novel chemical modifications demonstrated to improve siRNA specificity, Silencer Select siRNAs are highly potent with low off-target effects.
Bioinformatic filters applied in the Silencer Select algorithm

Although using siRNAs at low concentrations decreases off-target effects, further specificity gains can be made using bioinformatic filtering to predict and eliminate potentially inefficient siRNAs.
Our Silencer Select siRNA design process helps minimize off-target effects via:
- Stringent screening: identify and remove siRNAs with a high propensity for off-target effects by using a rigorous five-step process.
- Silencer Select toxicity classifier: helps eliminate sequences predicted to cause off-target apoptotic phenotypes to help ensure safer and more reliable siRNA performance.
- Seed region filtering: remove siRNAs with seed regions that resemble naturally occurring miRNAs and select those with the fewest seed region matches in the 3'UTRs of off-target transcripts to reduce miRNA pathway-related off-target effects.
The chemical modifications in Silencer Select siRNAs
Silencer Select siRNAs incorporate Locked Nucleic Acid (LNA) modifications to enhance their stability, binding affinity, and specificity. LNA-modified bases have the ribose ring chemically "locked" by a methylene bridge connecting the 2'-oxygen and the 4'-carbon. Adding LNA modifications to Silencer Select siRNA leads to:
- Consistent guide strand bias, which has been shown to correlate strongly with knockdown efficiency
- Prevention of the passenger strand from inducing silencing, which serves to reduce off-target effects (Figure 3)
- A reduced number of non-targeted, differentially expressed genes detected by gene expression array by up to 90% as compared to unmodified siRNAs (Figure 4)
- No negative impact on silencing efficiency and therefore do not compromise the expected on-target phenotypes

Figure 3. Silencer Select siRNA modifications reduce off-target effects and yield more reliable phenotypic data. Modified siRNAs preserved expected mitosis and apoptosis phenotypes for PLK and WEE1. In contrast the off-target apoptotic phenotypes elicited by 10 unmodified siRNAs were eliminated with addition of the Silencer Select modifications-demonstrating the modifications support generation of more reliable data.

Cleaner, more consistent data
siRNA mediated gene silencing enables the study of gene function with exceptional power and ease. However, currently available siRNAs for the same target do not always produce the same phenotype due to a combination of inconsistent silencing and sequence-specific off-target effects. Silencer Select siRNAs are designed with modifications that help overcome these issues, leading to more reliable data

Multiple publications confirm that higher siRNA concentrations lead to increased off-target effects [2,3]. This insight drove our efforts to design more potent siRNAs that can be used at lower concentrations, by significantly enhancing our siRNA design algorithms.
Silencer Select siRNAs
- Are up to 100X more potent than competitor siRNAs.
- Can be routinely transfected at concentrations of ≤5 nM and still retain their silencing power.
- Result in fewer off-target effects when used at these lower concentrations.
- Cost less per experiment than siRNAs used at higher concentrations.

Figure 5.Silencer Select siRNAs offer up to 100X higher potency compared to other siRNAs. Comparison of target knockdown in HeLa cells using Silencer Select siRNAs and siRNAs from two other suppliers to the same 10 different targets demonstrates the superior potency of Silencer Select siRNAs.
100% guarantee: One of the best guarantees in the industry
At Thermo Fisher Scientific, we guarantee that when you purchase two Invitrogen Silencer Select Pre-designed siRNA to the same target, those two siRNAs will silence the target mRNA by 70% or more. To qualify for the guarantee, siRNAs must have been transfected at ≥5 nM with proper controls and mRNA levels detected 48 hours post-transfection. Follow the link to learn more about our industry-leading guarantee.
Silencer Select validated siRNAs
A number of Invitrogen siRNAs targeting common human targets have been functionally tested by Thermo Fisher Scientific scientists and verified to reduce target mRNA levels by 70% or greater. The data sheet included with each of these siRNAs shows the extent of mRNA knockdown observed during testing and the exon targeted by the siRNA. We offer full sequences for predesigned and validated siRNAs with purchase.
Silencer Select siRNAs for long non-coding RNAs
Long non-coding RNAs (lncRNAs) are a type of non-coding RNAs (ncRNAs), typically over 200 nt, that are abundant in the mammalian transcriptome.
Features of Silencer Select siRNAs targeting lncRNAs
- Comprehensive portfolio of Silencer Select siRNAs targeting over 5,000 lncRNA targets
- Validated using databases including Gencode v27 for optimal coverage of lncRNA loci, pseudogene loci, and alternatively spliced transcripts
- Pre-designed siRNAs closely match existing TaqMan® non-coding assays
Tips for successful knockdown of lncRNA using Silencer Select siRNA
lncRNAs typically have lower expression levels compared to coding genes and expression can vary across cell lines. Improve results for knockdown of lncRNA with these suggestions:
- Verify expression levels in your cells of interest prior to your experiment.
- Choose at least three siRNAs for each lncRNA of interest.
- Use locked nucleic acid (LNA) modified siRNA (e.g., Silencer Select siRNA) over unmodified or earlier-generation siRNA for greater potency and lower off-target effects.
- Use a reliable transfection reagent such as Invitrogen Lipofectamine RNAiMAX Transfection Reagent for high transfection efficiency and improved cell viability.
Efficient knockdown of lncRNAs using Silencer Select siRNA
Results of targeted knockdown in HeLa and HEK293T cells indicate that two or more siRNA produced at least 50% knockdown. 80% knockdown or more was obtained for six out of the seven targets. Knockdown was effective for lncRNA with nuclear or cytoplasmic localization.
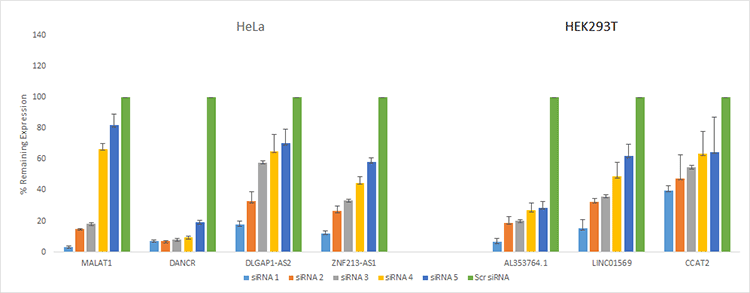
Figure 6. lncRNA produces effective knockdown across different targets. siRNA transfection of HeLa and HEK293T cells highlights the exceptional performance of Silencer Select lncRNA to knockdown expression across different gene targets.
Cells maintain high cell viability post siRNA transfection
Transfection reagents alone or in combination with siRNAs may lead to a nonspecific decrease in cell viability. This may impact interpretation of knockdown results. Five different siRNA against seven different lncRNA were transfected into the indicated cells. No significant decrease in viability was detected.
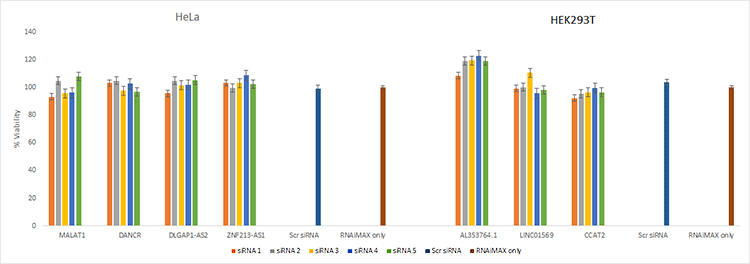
Figure 7. Cells transfected with siRNA retain high viability. Cell viability data demonstrates the low toxicity of different lncRNA transfected into HeLa and HEK293T cells.
Tips for successful knockdown of lncRNA using Silencer Select siRNA
lncRNAs typically have lower expression levels compared to coding genes and expression can vary across cell lines. Improve results for knockdown of lncRNA with these suggestions:
- Verify expression levels in your cells of interest prior to your experiment.
- Choose at least three siRNAs for each lncRNA of interest.
- Use locked nucleic acid (LNA) modified siRNA (e.g., Silencer Select siRNA) over unmodified or earlier-generation siRNA for greater potency and lower off-target effects.
- Use a reliable transfection reagent such as Invitrogen Lipofectamine RNAiMAX Transfection Reagent for high transfection efficiency and improved cell viability.
Efficient knockdown of lncRNAs using Silencer Select siRNA
Results of targeted knockdown in HeLa and HEK293T cells indicate that two or more siRNA produced at least 50% knockdown. 80% knockdown or more was obtained for six out of the seven targets. Knockdown was effective for lncRNA with nuclear or cytoplasmic localization.

Figure 6. lncRNA produces effective knockdown across different targets. siRNA transfection of HeLa and HEK293T cells highlights the exceptional performance of Silencer Select lncRNA to knockdown expression across different gene targets.
Cells maintain high cell viability post siRNA transfection
Transfection reagents alone or in combination with siRNAs may lead to a nonspecific decrease in cell viability. This may impact interpretation of knockdown results. Five different siRNA against seven different lncRNA were transfected into the indicated cells. No significant decrease in viability was detected.

Figure 7. Cells transfected with siRNA retain high viability. Cell viability data demonstrates the low toxicity of different lncRNA transfected into HeLa and HEK293T cells.
Silence Select RNA synthesis and turnaround times
We synthesize and purify each siRNA in an advanced facility that is ISO13485 and ISO9001 certified to meet the highest quality standards. As part of our rigorous quality control procedures, each RNA oligonucleotide is analyzed by LCMS and/or analytical HPLC. The result is premium quality siRNA that is purified and ready to use.
References
- Wang, X., Wang, X., Varma, RK, Beauchamp, L., Magdaleno, S, Sendera, TJ (2009) Selection of hyperfunctional siRNAs with improved potency and specificity. Nucleic Acids Res. 37(22):e152
- Semizarov D, Frost L, Sarthy A, Kroeger P, Halbert DN, Fesik SW (2003) Specificity of short interfering RNA determined through gene expression signatures. Proc Natl Acad Sci USA. 100(11):6347–6352.
- Jackson AL, Bartz SR, Schelter J, Kobayashi SV, Burchard J, Mao M, Li B, Cavet G, Linsley PS (2003) Expression profiling reveals off-target gene regulation by RNAi. Nat Biotechnol. 21(6):635–637.
We guarantee that when you purchase two Silencer Select Pre-designed siRNA to the same target, then those two siRNAs will silence the target mRNA by 70% or more. To qualify for the guarantee, siRNA's must have been transfected at ≥5 nM and mRNA levels detected 48h post transfection. Real-Time RT-PCR is recommended but not required for this application. Customers must also show > 80% knockdown with a positive control siRNA to demonstrate transfection efficiency. If the guaranteed level of knockdown is not observed and an appropriate positive control is successful, a onetime replacement of up to two new Silencer Select siRNA designs will be provided free of charge. This guarantee does not extend to any replacement product.
Reducing false positives and false negatives in siRNA experiments
(Open access to Ambion siRNA sequences and associated data)
In recent years, siRNA screens have highlighted issues with the technology stemming from off target effects, variable level of knockdown efficacy, and low-level confidence in hits from screening campaigns.
A recent genome-wide screen published in Nature by researchers at the National Institutes of Health highlights an effective approach to circumvent the challenges currently being faced by the screening community. New bioinformatics techniques to decipher real positives from false positives that arise from the seed region have been highlighted in this publication. Seed sequence driven off target effects have been extensively observed in siRNAs screens as these stem from the miRNA driven seed sequence. This seminal work highlights usage of high potency Silencer Select siRNA at lower siRNA concentration as arrayed singles allowing identification of off targets (false positive or false negative) stemming from the seed sequence. Such deconvolution is not possible for a pooled approach where unique biology can be observed owing to two or more siRNAs working in a concerted manner.
National Center for Advancing Translational Sciences (NCATS) and Life Technologies are also providing all researchers access to siRNA sequences and associated data on PubChem from Life Technologies’ Silencer Select siRNA library, which includes 65,000 siRNA sequences targeting more than 20,000 human genes.
Read the research article here
Get the siRNA sequences here
PRODUCT REPLACEMENT AS SET FORTH ABOVE IS LIFE TECHNOLOGIES’ SOLE OBLIGATION AND CUSTOMER’S SOLE AND EXCLUSIVE REMEDY WITH RESPECT TO THE GUARANTEE GIVEN ABOVE. To arrange a replacement, contact Technical Services. Please be ready to supply data indicating transfection efficiency and measurement of mRNA knockdown. Contact Technical Support if you require such a replacement.
Technical inquires:
Our Technical Application Scientists are available to help assist you at techsupport@thermofisher.com
Ordering & Order Status inquires:
If you have questions about pre-designed RNAi orders and order status, please contact us at genomicorders@thermofisher.com
If you have any questions about Custom RNAi orders and order status, please contact us at RNAiSupport@thermofisher.com
For Research Use Only. Not for use in diagnostic procedures.
