Search
Active site probes are powerful tools used to label and enrich proteins based on their enzymatic activity, allowing researchers to isolate only the functional forms of target enzymes. These kits enable activity-based protein profiling for a wide range of applications in drug discovery and disease research.
An active site in an enzyme is the specific region where a substrate binds and a chemical reaction occurs. For example, in kinases, the ATP-binding pocket is a well-studied active site that drives phosphorylation reactions. Active sites are determined experimentally using mutagenesis, binding assays or structural biology techniques and active site probes simplify this process by selectively labeling functional residues. This makes it possible to map and study active sites directly in complex samples. In practice, active site probes are used to profile enzyme activity, assess inhibitor selectivity and enrich for functional proteins rather than inactive or denatured forms. For more options and to explore additional protein enrichment tools and kits, visit our Protein Enrichment solutions page.
How active site probes are used
Active site probes enable functional enzyme profiling and small-molecule inhibitor screening across complex biological samples. Active site probes do the following:
- Screen small-molecule inhibitors in cell lysates, subcellular fractions, tissues, and recombinant proteins
- Enrich and profile nucleotide-binding proteins, including kinases and G-proteins
- Perform dose-dependent profiling of small molecules
- Identify enzyme targets and off-targets at scale
- Detect and quantify labeled enzymes by western blot, fluorescent gel imaging, or mass spectrometry
- Enable high-throughput mass spectrometry analysis of hundreds of enzymes in a single experiment
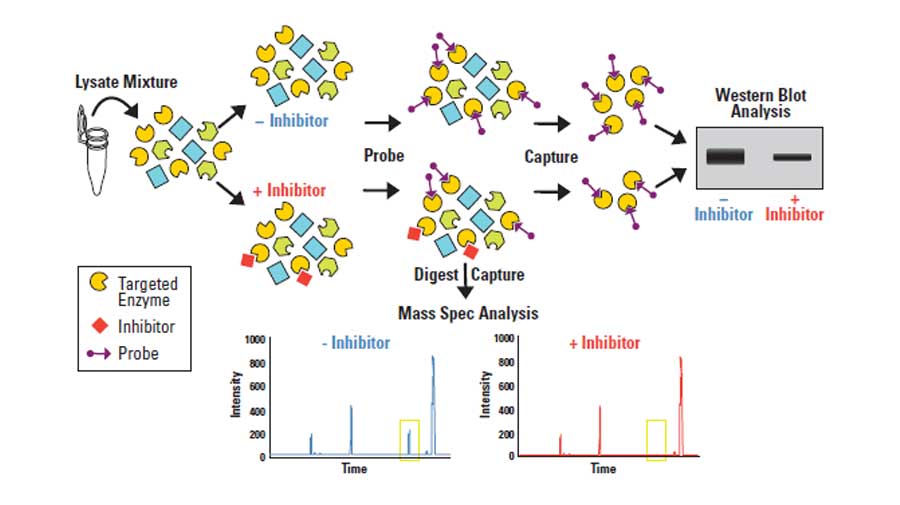
Features of Thermo Scientific products that use ActivX™ probes
Thermo Scientific active site probes covalently bind to the active sites of kinases, GTPases and serine hydrolases to enable selective enrichment and profiling of target enzyme classes. The structure of the probes consists of an active site-specific reactive group, a linker region and either desthiobiotin or TAMRA tags for enrichment or detection. Active site reactive groups are typically electrophilic compounds which covalently link to nucleophilic resides in enzyme active sites. All active site probes can be used to determine inhibition of enzymes by small molecules, and some probes also preferentially react with only active enzymes, allowing for activity-based proteomic profiling (ABPP).
These products are based on Activity-Based Protein Profiling Technology, licensed from ActivX Biosciences, Inc.
Kinase active site probes and enzyme enrichment kits
One class of active site probes is based on derivatives of ATP, ADP or GTP nucleotides. ATP- and ADP-based probes covalently modify the active site of ATPases including kinases, chaperones and metabolic enzymes. The GTP probe specifically labels small GTPases and G-protein coupled receptor GTPase subunits. As many kinases, GTPases, and other nucleotide binding proteins bind nucleotides or inhibitors even when they are enzymatically inactive, these reagents allow both inactive and active enzymes to be profiled in a complex sample. Preincubation of samples with small molecule inhibitors that compete for active site probes can be used to determine inhibitor binding affinity. In addition, active site directed nucleotide probes can be used to determine inhibitors off targets.
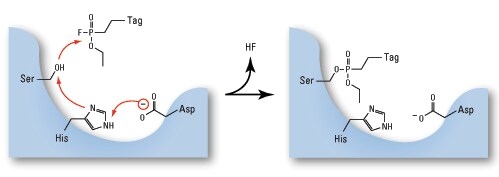
Mechanism and chemical structures of Thermo Scientific Active Site Probes for kinases, other ATPases, and GTPases. (A) Nucleotide analogues bind to the active sites of ATPases or GTPases and the biotin affinity tag is irreversibly transferred to highly conserved lysine residues in the active site. (B) Structures of desthiobiotin nucleotide analogues. Desthiobiotin binding to streptavidin is easily reversible under acidic elution conditions, allowing high recovery of labeled proteins and peptides. Desthiobiotin is attached to the nucleotide through a labile acyl phosphate linkage, allowing efficient desthiobiotin label transfer to amines near the active site. ATP and ADP nucleotide analogues label a complementary set of ATPases, which is likely due to differences in the proximity of the acyl phosphate linkage to conserved lysines near the active site.
View products
Serine hydrolase active site probes
Fluorophosphonate (FP) probes are another class of active site probes which are specific for active serine hydrolases. Since these reagents only label active enzymes, they are able to monitor enzymatic activity in addition to being used for studying inhibitor binding affinities. The serine hydrolase family includes a variety of enzymes such as cholinesterases, hydrolases, lipases, and proteases like trypsin, kallikreins and dipeptidyl peptidases. Since many of these enzymes are expressed as inactive pro-proteins, the ability of these probes to assess activity is especially advantageous over traditional protein or RNA expression profiling techniques which only measure abundance.
A.
B.
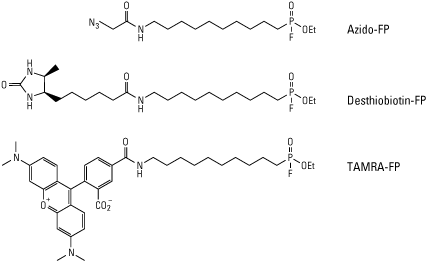
View products
仅供科研使用,不可用于诊断目的。

